Pediatric incubation time for COVID-19 using CORD-19 data
This post replicates my recently-posted Kaggle notebook using the the CORD-19 dataset which has more 37K full-text COVID-related articles. The goal of post is to show how to filter articles that are look for incubation period of the disease with the goal of finding a subset of articles that have a pediatric reference.
Background
A wide variety of estimates of the incubation time have been seen in the rapidly evolving Covid-19 literature. As of March 27th, 2020 the most common number reported in the popular press appears to be the 5.2 day estimate. While the variation in point estimates continues to be substantial, an average range of 5-6 days appears to be appropriate (for the mean). There has also been a limited discussion on the possibility of age-based differences in the incubation period. This kernel will go through the CORD-19 dataset and extract relevant sentences in regards to different moments of the the incubation period (mean, median, lower-bound, and upper-bound). At the end of the notebook, a subset of relevant papers that mention pediatric populations will be explored. As of now, there does not appear to be strong evidence for age related-difference in incubation periods.
Several technical notes.
- The
df_txt.csvload in the script below was generated with a similar method to xhlulu’s kernel. - A utility script is being used to help with the parsing. This can be found on githib
- After the relevant sentences are found, a function
record_valsis used to allow the user to manually select the sentences with “y(es)/n(o)” - I manually annotated the moments in the sentences
import numpy as np
import pandas as pd
import os
import re
import seaborn as sns
from datetime import datetime as dt
from support_funs_incubation import stopifnot, uwords, idx_find, find_beside, ljoin, sentence_find, record_vals, color_printer
# load data
df = pd.read_csv('df_txt.csv')
df['date'] = pd.to_datetime(df.date)
print(df.shape)
# remove prefix from some abstracts: publically funded repositories.... etc
pref = 'COVID-19 resource centre remains active.'
for ii, aa in enumerate(df.abstract):
if isinstance(aa, float): # nan
continue
hit = re.search(pref, aa)
if hit:
df.abstract.iloc[ii] = aa[hit.span()[1] + 1:]
(27936, 6)
Section 1: Summary statistics
The code block below will calculate the number of ‘covid’ and ‘nCoV’ mentions in the texts and abstracts of the corpus. The first journal articles referencing ‘covid’ and ‘nCoV’ start on January 1st, 2020. The last relevant articles are around a week old as of March 27, 2020. The majority of articles use either ‘Covid-2019’ or ‘2019-nCoV’, although there are some exceptions.
# Find ways in which covid and ncov are referred to
regex_ncov = r'(20)?19(\-)?ncov|ncov(\-)?(20)?19'
regex_covid = r'covid(\-)?(20)?19'
# row indices
idx_covid_abs = np.where(idx_find(df.abstract, regex_covid))[0]
idx_ncov_abs = np.where(idx_find(df.abstract, regex_ncov))[0]
idx_union_abs = np.union1d(idx_covid_abs, idx_ncov_abs)
di_regex = {'covid': regex_covid, 'ncov': regex_ncov}
di_idx = {'covid': idx_covid_abs, 'ncov': idx_ncov_abs}
print('%i possible "covid" articles (using abstract)\n'
'%i possible nCoV articles (using abstract)\n'
'Union: %i, interection: %i' %
(len(idx_covid_abs), len(idx_ncov_abs), len(idx_union_abs),
len(np.intersect1d(idx_covid_abs, idx_ncov_abs))))
dfmt = '%B %d, %Y'
date_ncov_min = df.date.iloc[idx_ncov_abs].min().strftime(dfmt)
date_ncov_max = df.date.iloc[idx_ncov_abs].max().strftime(dfmt)
date_covid_min = df.date.iloc[idx_covid_abs].min().strftime(dfmt)
date_covid_max = df.date.iloc[idx_covid_abs].max().strftime(dfmt)
print('First and last nCoV article: %s & %s\n'
'First and last covid-19 article: %s & %s' %
(date_ncov_min, date_ncov_max, date_covid_min, date_covid_max))
holder = []
for term in di_regex:
regex = di_regex[term]
idx = di_idx[term]
dat_abstract = uwords(df.abstract.iloc[idx], regex).assign(doc='abstract')
dat_txt = uwords(df.txt.iloc[idx], regex).assign(doc='txt')
dat = pd.concat([dat_abstract, dat_txt])
dat = dat.groupby('term').n.sum().reset_index()
dat.insert(0, 'tt', term)
holder.append(dat)
df_term = pd.concat(holder).reset_index(drop=True)
# Term usage
print(df_term)
425 possible "covid" articles (using abstract)
229 possible nCoV articles (using abstract)
Union: 610, interection: 44
First and last nCoV article: January 01, 2020 & March 17, 2020
First and last covid-19 article: January 01, 2020 & March 20, 2020
tt term n
0 covid covid-19 6612
1 covid covid-2019 68
2 covid covid19 93
3 ncov 2019-ncov 3733
4 ncov 2019ncov 48
5 ncov ncov-2019 49
6 ncov ncov2019 3
Section 2: Incubation period
To see how ‘incubation’ is being used in the corpus, it is useful to see the preceeding and succeeding word. While ‘incubation period’ is the most common expression, others such as ‘incubation time’ or ‘incubation period’ are in use too.
pat_peds = r'infant|child|pediatric|age\<'
idx_incubation = []
idx_peds = []
for ii in idx_union_abs:
abs, txt = df.abstract[ii], df.txt[ii]
corpus = abs + '. ' + txt
if re.search(r'incubation', corpus, re.IGNORECASE) is not None:
idx_incubation.append(ii)
if re.search(pat_peds, corpus, re.IGNORECASE) is not None:
idx_peds.append(ii)
idx_incubation_peds = np.intersect1d(idx_incubation, idx_peds)
print('%i incubation articles, with %i pediatric articles, %i overlap' %
(len(idx_incubation), len(idx_peds), len(idx_incubation_peds)))
# What is the most common word to appear before/after incubation?
holder_l, holder_r = [], []
for ii in idx_incubation:
abs, txt = df.abstract[ii], df.txt[ii]
corpus = abs + '. ' + txt
rterm = find_beside(corpus, 'incubation', tt='right')
lterm = find_beside(corpus, 'incubation', tt='left')
holder_r.append(rterm)
holder_l.append(lterm)
dat_suffix = pd.Series(ljoin(holder_r)).str.lower().value_counts().reset_index().rename(
columns={0: 'n', 'index': 'suffix'})
dat_prefix = pd.Series(ljoin(holder_l)).str.lower().value_counts().reset_index().rename(
columns={0: 'n', 'index': 'prefix'})
print(pd.concat([dat_suffix.head(20),dat_prefix.head(20)],1))
suffix = ['period', 'time', 'distribution', 'duration', 'interval', 'rate', 'mean', 'median', 'estimation']
suffix = [z + r'(s)?' for z in suffix]
pat_incubation = [r'incubation\s'+z for z in suffix]
194 incubation articles, with 74 pediatric articles, 40 overlap
suffix n prefix n
0 period 822 the 473
1 periods 75 mean 84
2 time 50 of 59
3 of 16 median 44
4 and 9 an 26
5 distribution 8 estimated 25
6 with 8 their 22
7 times 6 long 21
8 duration 5 average 20
9 at 5 longer 19
10 to 3 and 17
11 interval 3 maximum 13
12 was 3 shorter 10
13 individual 3 day 10
14 for 3 with 10
15 sample 3 as 9
16 in 3 asymptomatic 8
17 using 2 in 6
18 bypass 2 infectious 6
19 patients 2 different 6
Section 4: Manual curation
Now that a total of 194 articles have been found with relevant sentences, a manual curation will be performed to select which sentences are relevant and allow the user to annotate the data with the stated moments. Sentences were selected if they estimated an incubation period from actual data rather than used existing estimates.
# Run sentence finder if output does not exist
do_run = 'sentence_flag.csv' not in os.listdir()
if do_run:
keepers = []
for jj, ii in enumerate(idx_incubation):
abs, txt = df.abstract[ii], df.txt[ii]
corpus = abs + '. ' + txt
idx_sentences = sentence_find(corpus, pat_incubation)
if len(idx_sentences) > 0:
try:
dd = df.loc[ii,'date'].strftime('%B %d, %Y')
except:
dd = 'NaN'
print('</p><p>----- Title: %s, date: %s, index: %i (%i of %i) ----' %
(df.loc[ii, 'title'], dd , ii,jj+1,len(idx_incubation)))
tmp = record_vals(idx_sentences)
dat = pd.DataFrame(tmp,columns=['pos','txt']).assign(idx = ii)
keepers.append(dat)
dat_sentences = pd.concat(keepers)
dat_sentences = dat_sentences[['idx','pos','txt']]
dat_sentences['txt'] = dat_sentences.txt.str.replace('\n','')
dat_sentences = df.iloc[idx_incubation][['source','title','doi','date']].rename_axis('idx').reset_index().merge(
dat_sentences,on='idx',how='right')
dat_sentences.to_csv('sentence_flag.csv',index=False)
Section 5: Analyze moments of incubation period
Load the manually annotated data with the added moments column.
df_moments = pd.read_csv('sentence_flag.csv')
df_txt = df_moments[['title','pos','txt']].copy()
df_moments.drop(columns = ['pos','txt'],inplace=True)
df_moments['date'] = pd.to_datetime(df_moments.date)
moments = df_moments.moments.str.split('\;',expand=True).reset_index().melt('index')
moments = moments[moments.value.notnull()].reset_index(drop=True).drop(columns='variable')
tmp = moments.value.str.split('\=',expand=True)
moments = moments.drop(columns='value').assign(moment=tmp.iloc[:,0], val=tmp.iloc[:,1].astype(float))
df_moments = df_moments.drop(columns='moments').reset_index().merge(moments,on='index',how='right').drop(columns='index')
# Print off key sentences
print('A total of %i unique studies' % (df_moments.title.unique().shape[0]) )
print('\n\n')
for ii, rr in df_txt.iterrows():
print('</p><p>----- Article: %s -----' % rr['title'] )
idx = [int(z) for z in re.findall(r'\d+', rr['pos'])]
idx = np.array(idx).reshape([int(len(idx) / 2), 2])
idx = [tuple(idx[i]) for i in range(idx.shape[0])]
sentence = rr['txt']
idx_sentence = (idx,sentence)
color_printer(idx_sentence)
print('\n')
A total of 30 unique studies
----- Article: Incubation Period and Other Epidemiological Characteristics of 2019 Novel Coronavirus Infections with Right Truncation: A Statistical Analysis of Publicly Available Case Data ----- Our results show that the incubation period falls within the range of 2–14 days with 95% confidence and has a mean of around 5 days when approximated using the best-fit lognormal distribution.
----- Article: Incubation Period and Other Epidemiological Characteristics of 2019 Novel Coronavirus Infections with Right Truncation: A Statistical Analysis of Publicly Available Case Data ----- The mean incubation period was estimated at 5.0 days (95% credible interval [CI]: 4.2, 6.0) when excluding Wuhan residents (n = 52) and 5.6 days (95% CI: 5.0, 6.3) when including Wuhan residents (n = 158).
----- Article: 2019-nCoV (Wuhan virus), a novel Coronavirus: Human-to-human transmission, travel-related cases, and vaccine readiness ----- The mean incubation period was estimated as 4.6 days with a range of 2 to 8 days between symptom onset and hospitalization.
----- Article: The outbreak of COVID-19: An overview ----- COVID-19 has a mean incubation period of 5.2 days (95% confidence interval, 4.1-7.0).
----- Article: Clinical findings in a group of patients infected with the 2019 novel coronavirus (SARS-Cov-2) outside of Wuhan, China: retrospective case series ----- Among 56 patients who could provide the exact date of close contact with someone with confirmed or suspected SARS-Cov-2 infection, the median incubation period from exposure to symptoms was 4 days (interquartile range 3-5 days).
----- Article: COVID-19 outbreak on the Diamond Princess cruise ship: estimating the epidemic potential and effectiveness of public health countermeasures ----- 8 In the situation of no removal (ill persons taken off the ship to be isolated in a Japanese hospital), the incubation period (or, the latent period), was estimated to be approximately 5 days (ranging from 2 to 14 days).
----- Article: Comparative genomic analysis revealed specific mutation pattern between human coronavirus SARS-CoV-2 and Bat-SARSr-CoV RaTG13 ----- SARS-CoV-2 has a similar incubation period (median, 3.0 days) and a relatively lower fatality rate than SARS-CoV or MERS-CoV (1), but it is estimated that the reproductive number of SARS-CoV-2 is higher than that of SARS-CoV (2) .
----- Article: The incubation period of 2019-nCoV infections among travellers from Wuhan, China ----- Using the travel history and symptom onset of 88 confirmed cases that were detected outside Wuhan, we estimate the mean incubation period to be 6.4 (5.6 - 7.7, 95% CI) days, ranging from 2.1 to 11.1 days (2.5th to 97.5th percentile).
----- Article: The incubation period of 2019-nCoV from publicly reported confirmed cases: estimation and application ----- Here, we use data from public reports of 101 confirmed cases in 38 provinces, regions, and countries outside of Wuhan (Hubei province, China) with identifiable exposure windows and known dates of symptom onset to estimate the incubation period of 2019-nCoV. We estimate the median incubation period of 2019-nCoV to be 5.2 days (95% CI: 4.4, 6.0), and 97.5% of those who develop symptoms will do so within 10.5 days (95% CI: 7.3, 15.3) of infection.
----- Article: The incubation period of 2019-nCoV from publicly reported confirmed cases: estimation and application ----- The estimated mean incubation period was 5.5 days.
----- Article: The incubation period of 2019-nCoV from publicly reported confirmed cases: estimation and application ----- Based on cases detected inside mainland China (n=29), the median incubation period is 4.6 days (95% CI: 3.4, 6.0), with a 95% range of 2.7 (95% CI: 1.2, 4.6) to 7.9 (95% CI: 3.9, 13.1) days.
----- Article: Clinical characteristics of 2019 novel coronavirus infection in China ----- The median incubation period was 3.0 days (range, 0 to 24.0 days).
----- Article: The cross-sectional study of hospitalized coronavirus disease 2019 patients in Xiangyang, Hubei province ----- Incubation time ranged from one to twenty days with a mean period of 8.09 days (SD 4.99).
----- Article: A descriptive study of the impact of diseases control and prevention on the epidemics dynamics and clinical features of SARS-CoV-2 outbreak in Shanghai, lessons learned for metropolis epidemics prevention ----- The mean incubation period is 6.4 days (95% 175 CI 5.3 to 7.6) and the 5th and 95th percentile of the distribution was 0.97 and 13.10, 176 respectively ( Figure 3 -A).
----- Article: Epidemiological characteristics of 1212 COVID-19 patients in Henan, China ----- The following findings are obtained: 1) COVID-19 patients in Henan show gender (55% vs 45%) and age (81% aged between 21 and 60) preferences, possible causes were explored; 2) Statistical analysis on 483 patients reveals that the estimated average, mode and median incubation periods are 7.4, 4 and 7 days; Incubation periods of 92% patients were no more than 14 days; 3) The epidemic of COVID-19 in Henan has undergone three stages and showed high correlations with the numbers of patients that recently return from Wuhan; 4) Network analysis on the aggregate outbreak phenomena of COVID-19 revealed that 208 cases were clustering infected, and various people's Hospital are the main force in treating patients.
----- Article: Evolving epidemiology of novel coronavirus diseases 2019 and possible interruption of local transmission outside Hubei Province in China: a descriptive and modeling study ----- We estimated a mean incubation period of 5.2 days (95%CI:1.8-12.4), with the 95th percentile of the distribution at 10.5 days.
----- Article: Epidemiologic and Clinical Characteristics of 91 Hospitalized Patients with COVID-19 in Zhejiang, China: A retrospective, multi-centre case series ----- The median of incubation period was 6 (IQR, 3-8) days and the median time from first visit to a doctor to confirmed diagnosis was 1 (1-2) days.
----- Article: Estimate the incubation period of coronavirus 2019 (COVID-19) ----- Our results show the incubation mean and median of COVID-19 are 5.84 and 5.0 days respectively and there is a statistical significance with the role of gender.
----- Article: Prevalence and clinical features of 2019 novel coronavirus disease (COVID-19) in the Fever Clinic of a teaching hospital in Beijing: a single-center, retrospective study ----- An incubation period was elicited from 12 patients (57.1%), ranging from 2 to 10 days with a median of 6.5 days.
----- Article: Transmission characteristics of the COVID-19 outbreak in China: a study driven by data ----- The mean generation time is 3.3 days and the mean incubation period is 7.2 days.
----- Article: Transmission characteristics of the COVID-19 outbreak in China: a study driven by data ----- The best fit incubation period distribution is a gamma distribution with a mean 7.2 (6.8, 7.6) days and a variance 16.9 (14.0, 20.2), which correspond to a shape parameter 3.07 (2.62, 3.56) and a scale parameter 2.35 (2.00, 2.75).
----- Article: Analysis of epidemiological characteristics of coronavirus 2019 infection and preventive measures in Shenzhen China: a heavy population city ----- Taking account of the incubation period (mostly 3-7 days, with mean of 3.7 days) and the time between symptom onset and confirm of the diagnosis (6 day on average) [9, 12] , the peak of new confirmed cases coincided with the implementation of serial preventive strategy and measures,indicating these preventive strategies and measures were effective in preventing transmission of COVID-19 in Shenzhen.
----- Article: Epidemiologic Characteristics of COVID-19 in Guizhou, China ----- The median incubation period was 8.2 days (95% CI: 155 7.9 -9.5) in our study, which is longer than a recent report of 425 patients (8.2 days 156 vs. 5.2 days), this may be a results of recall bias, during epidemiological investigation, symptoms does not yet infect others, which assumes that all index cases should show 165 symptoms before their secondary cases.
----- Article: The effect of human mobility and control measures on the COVID-19 epidemic in China ----- Using detailed information on 38 cases for whom both the dates of entry to and exit from Wuhan are known, we estimate the mean incubation period to be 5.1 days (std.
----- Article: Transmission interval estimates suggest pre-symptomatic spread of COVID-19 ----- Results: The mean incubation period was 7.1 (6.13, 8.25) days for Singapore and 9 (7.92, 10.2)days for Tianjin.
----- Article: Transmission interval estimates suggest pre-symptomatic spread of COVID-19 ----- The estimated median incubation period for pre-Jan 31 cases in Tianjin is 6.9 days; the q = (0.025, 0.975) quantiles are (2, 12.7) days.
----- Article: Transmission interval estimates suggest pre-symptomatic spread of COVID-19 ----- The estimated median incubation time is 5.46, with (0.025, 0.975) quantiles of (1.34, 11.1) days for early cases and 7.27 days (quantiles (1.31, 17.
----- Article: Transmission interval estimates suggest pre-symptomatic spread of COVID-19 ----- In the Singapore dataset, we find that the median incubation period is 6.55 days with the Weibull
----- Article: Transmission and clinical characteristics of coronavirus disease 2019 in 104 outside-Wuhan patients, China ----- The median incubation period was 6 (rang, 1-32) days, of 8 patients ranged from 18 to 32 days.
----- Article: Transmission of corona virus disease 2019 during the incubation period may lead to a quarantine loophole ----- Results: The estimated mean incubation period for COVID-19 was 4.9 days (95% confidence interval [CI], 4.4 to 5.4) days, ranging from 0.8 to 11.1 days (2.5th to 97.5th percentile).
----- Article: Estimation of incubation period distribution of COVID-19 using disease onset forward time: a novel cross-sectional and forward follow-up study ----- The estimated median of incubation period is 8.13 days (95% confidence interval [CI]: 7.37-8.91), the mean is 8.62 days (95% CI: 8.02-9.28), the 90th percentile is 14.65 days (95% CI: 14.00-15.26), and the 99th percentile is 20.59 days (95% CI: 19.47, 21.62).
----- Article: Clinical Characteristics of 34 Children with Coronavirus Disease-2019 in the West of China: a Multiple-center Case Series ----- In accordance with the features of indirect contact exposure, the incubation period was 10.50 (7.75 -25.25) days in paediatric patients, while 4.00 (2.00 -7.00) days of incubation was revealed for patients in all age groups [1] .
----- Article: Early Epidemiological and Clinical Characteristics of 28 Cases of Coronavirus Disease in South Korea ----- The incubation period was estimated to be 4.1 days based on the date of symptom onset and first exposure (among 9 patients, excluding 1 patient with an unclear date of onset) shown in Figure 1 and Table 2 .
----- Article: Early Epidemiological and Clinical Characteristics of 28 Cases of Coronavirus Disease in South Korea ----- The estimated incubation period was 4.6 days to develop COVID-19, which was shorter than the period of 5.2 days reported in China [5] .
----- Article: Investigation of three clusters of COVID-19 in Singapore: implications for surveillance and response measures ----- The median incubation period of SARS-CoV-2 was 4 days (IQR 3–6).
----- Article: Early epidemiological analysis of the coronavirus disease 2019 outbreak based on crowdsourced data: a population-level observational study ----- On the basis of 33 patients with a travel history to Wuhan, we estimated the median incubation period for COVID-19 to be 4·5 days (IQR 3·0-5·5; appendix p 2).
----- Article: Characteristics of COVID-19 infection in Beijing ----- The median incubation period was 6.7 days, the interval time from between illness onset and seeing a doctor was 4.5 days.
----- Article: Epidemiological and clinical features of COVID-19 patients with and without pneumonia in Beijing, China ----- The mean incubation period of COVID-19 was 8.42 days
di_moments = {'lb':'Lower-bound','ub':'Upper-bound','mu':'Mean','med':'Median',
'q2':'25th percentile','q3':'75th percentile'}
# Plot the moments over time
g = sns.FacetGrid(data=df_moments.assign(moment=lambda x: x.moment.map(di_moments)),
col='moment',col_wrap=3,sharex=True,sharey=False,height=4,aspect=1)
g.map(sns.lineplot,'date','val',ci=None)
g.map(sns.scatterplot,'date','val')
g.set_xlabels('');g.set_ylabels('Days')
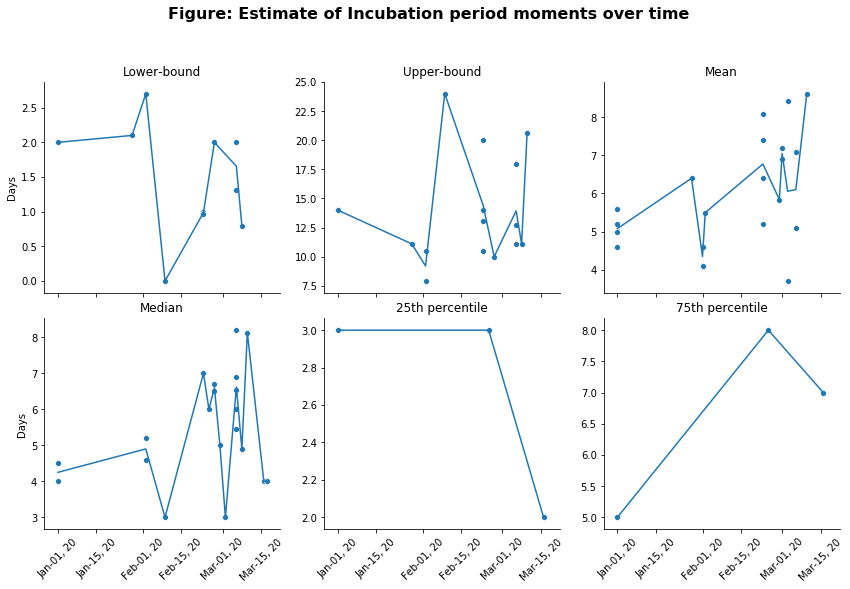
g.fig.suptitle(t='Figure: Estimate of Incubation period moments over time',size=16,weight='bold')
g.fig.subplots_adjust(top=0.85)
for ax in g.axes.flat:
ax.set_title(ax.title._text.replace('moment = ', ''))
# dates = [dt.strftime(dt.strptime(z,'%Y-%m-%d'),'%b-%d, %y') for z in dates]
xticks = [737425., 737439., 737456., 737470., 737485., 737499.]
lbls = ['Jan-01, 20', 'Jan-15, 20', 'Feb-01, 20', 'Feb-15, 20', 'Mar-01, 20', 'Mar-15, 20']
g.set_xticklabels(rotation=45,labels=lbls)
g.set(xticks = xticks)
ave = df_moments.groupby('moment').val.mean().reset_index().rename(columns={'moment':'Moment','val':'Average'}).assign(Moment=lambda x: x.Moment.map(di_moments))
print(np.round(ave,1))
Moment Average
0 Lower-bound 1.5
1 Median 5.5
2 Mean 6.0
3 25th percentile 2.8
4 75th percentile 6.2
5 Upper-bound 13.9
The figure above shows thats the point estimates, especially for the mean, are quite noisy and range from just below 3 days, to just above 8 days.
Section 6: Pediatric references
Using the 30 articles found above, we can now see which papers might shed any clues on the incubation period for pediatric populations.
# Get the index
df_match = df_txt.merge(df,on='title',how='left').rename(columns={'txt_x':'sentence','txt_y':'txt_full'})
for jj, rr in df_match.iterrows():
try:
dd = rr['date'].strftime('%B %d, %Y')
except:
dd = 'NaN'
corpus = rr['abstract'] + '. ' + rr['txt_full']
peds_sentences = sentence_find(corpus, pat_peds)
incubation_sentences = sentence_find(corpus, pat_incubation)
if len(peds_sentences) > 0 and len(incubation_sentences) > 0:
print('</p><p>----- Title: %s, date: %s (%i of %i) ----' %
(rr['title'], dd, jj+1, df_match.shape[0]))
for ii_ss in peds_sentences + incubation_sentences:
color_printer(ii_ss)
print('\n')
----- Title: 2019-nCoV (Wuhan virus), a novel Coronavirus: Human-to-human transmission, travel-related cases, and vaccine readiness, date: January 01, 2020 (3 of 38) ---- As with the SARS-CoV, infections in children appear to be rare. No fatalities were reported in young children and adolescents and fatal disease was reported in 6.8% of patients < 60 years of age. The mean incubation period was estimated as 4.6 days with a range of 2 to 8 days between symptom onset and hospitalization. Early symptoms of MERS include fever, chills, cough, shortness of breath, myalgia, and malaise following a mean incubation period of 5 days, with a range of 2 to 13 days [23] .
----- Title: Clinical findings in a group of patients infected with the 2019 novel coronavirus (SARS-Cov-2) outside of Wuhan, China: retrospective case series, date: January 01, 2020 (5 of 38) ---- Most of the infected individuals in Zhejiang province were male patients, but the age range of patients is large as SARS-Cov-2 also infected children and those older than 65 years. Data were collected from 10 January 2020 to 26 January 2020.Main outcome measures Clinical data, collected using a standardised case report form, such as temperature, history of exposure, incubation period. The incubation period was defined as the time from exposure to the onset of illness, which was estimated among patients who could provide the exact date of close contact with individuals from Wuhan with confirmed or suspected SARS-Cov-2 infection. Among 56 patients who could provide the exact date of close contact with someone with confirmed or suspected SARS-Cov-2 infection, the median incubation period from exposure to symptoms was 4 days (interquartile range 3-5 days). Among the 33 patients who had symptoms for more than 10 days after illness onset, the median incubation period from exposure to symptoms was 3 days (interquartile range 3-4 days). It is possible that an even greater number of infected patients exist without a diagnosis because their symptoms were less severe and because of the incubation period.
----- Title: Epidemiologic and Clinical Characteristics of 91 Hospitalized Patients with COVID-19 in Zhejiang, China: A retrospective, multi-centre case series, date: February 25, 2020 (17 of 38) ---- There was 1 child (5 years . The median of incubation period was 6 (IQR, 3-8) days and the median time from first visit to a doctor to confirmed diagnosis was 1 (1-2) days. 11 They reported that the median age was 47.0 years, 41.90% were females, 31.30% had been to Wuhan and 71.80% had contacted people from Wuhan, and the average incubation period was 3.0 days. 13,14 The incubation period was defined as the time from the exposure to the confirmed or suspected transmission source to the onset of illness. The median of incubation period is 6 (IQR, 3-8) days, and number of days from first visit to a doctor till the case is confirmed is 1 (1-2). The median of incubation period was 6 (IQR, [3] [4] [5] [6] [7] [8] days and from first visit to a doctor to confirmed diagnosis was only 1 (1-2) days. 20 It appears that transmission is possible during the incubation period, and the carrier cannot be spotted.
----- Title: Estimate the incubation period of coronavirus 2019 (COVID-19), date: February 29, 2020 (18 of 38) ---- We found that the incubation periods of the groups with age>=40 years and age<40 years demonstrated a statistically significant difference. However, the incubation periods of the groups with age>=40 years and age<40 years show a statistically significant difference. To verify whether the incubation of the age>=40 group is different from that of the age<40 group statistically, Figure 3 compares All rights reserved. The Mann-Whitney rank test shows that there's a statistically significant difference between the incubation of age<40 and age>=40 groups with the null hypothesis: the medians of incubation period between two groups are the same. The p-values for corresponding alternative hypotheses: the age<40 group has a smaller incubation median is 0.00474. It suggests that the age<40 group has a shorter incubation period than the age>=40 group. Data is partitioned as the age>=40 and age<40 groups in visualization. However, we also find that the incubation data will no longer demonstrate the linear separability property when we partition it as age>=50 and age<50 groups or age>=55 All rights reserved. https://doi.org/10.1101/2020.02.24.20027474 doi: medRxiv preprint and age<55 groups. Our studies indicate that incubation periods of the age>=40 years and age<40 years groups not only statistically significant but also linearly separable in machine learning. Furthermore, different quarantine time should be applied to the age>=40 years and age<40 years groups for their different incubation periods. Accurate estimation of the incubation period of the coronavirus is essential to the prevention and control. However, it remains unclear about its exact incubation period though it is believed that symptoms of COVID-19 can appear in as few as 2 days or as long as 14 or even more after exposure. The accurate incubation period calculation requires original chain-of-infection data that may not be fully available in the Wuhan regions. In this study, we aim to accurately calculate the incubation period of COVID-19 by taking advantage of the chain-of-infection data, which is well-documented and epidemiologically informative, outside the Wuhan regions. To achieve the accurate calculation of the incubation period, we only involved the officially confirmed cases with a clear history of exposure and time of onset. Result: The incubation period of COVID-19 did not follow general incubation distributions such as lognormal, Weibull, and Gamma distributions. Result: The incubation period of COVID-19 did not follow general incubation distributions such as lognormal, Weibull, and Gamma distributions. We found that the incubation periods of the groups with age>=40 years and age<40 years demonstrated a statistically significant difference. The former group had a longer incubation period and a larger variance than the latter. It further suggested that different quarantine time should be applied to the groups for their different incubation periods. It is essential to know the accurate incubation period of COVID-19 for the sake of deciphering dynamics of its spread. The incubation period is the time from infection to the onset of the disease. Different viruses have different incubation periods that determine their different dynamics epidemiologically. The incubation period of H7N9 (Human Avian Influenza A) is about 6.5 days, but the incubation period for SARS-CoV is typically 2 to 7 days [5] [6] . However, it remains unclear about its exact incubation period of COVID-19, although WHO estimates it is between 2 to 14 days after exposure [8] . https://doi.org/10.1101/2020.02.24.20027474 doi: medRxiv preprint incubation period of COVID-19 by using original chain-of-infection data that may not be fully available in the Wuhan regions. Furthermore, it is also unknown whether the incubation time will show some statistically significant with respect to Age and Gender. In this study, we aim to accurately estimate the incubation period of COVID-19 by taking advantage of datasets with a well-documented history of exposure. Our results show the incubation mean and median of COVID-19 are 5.84 and 5.0 days respectively and there is a statistical significance with the role of gender. However, the incubation periods of the groups with age>=40 years and age<40 years show a statistically significant difference. For those cases whose incubation periods locate in an interval [ ! , " ], $# " " to represent its incubation period. The incubation will be calculated as = We propose a Monte Carlo simulation approach that takes advantage of bootstrap techniques to estimate incubation median and mean estimation for the small sample with 59 cases. It indicates that the incubation period median and mean of patients more than 40 years old are greater than those of patients less than 40 years old. https://doi.org/10.1101/2020.02.24.20027474 doi: medRxiv preprint The incubation of COVID-19 is not subject to neither of the widely used incubation distributions such as normal, lognormal, Gamma, and Weibull distributions well [12] . Although we can't reject Gamma distributions for its boundary line p-value (0.06086), it can be risky to use it to fit and estimate the distribution of the incubation period under a small sample size. It suggests that that the incubation period is somewhat correlated with age though not that strong. https://doi.org/10.1101/2020.02.24.20027474 doi: medRxiv preprint their incubation periods groups using different visualization tools. It indicates that the younger group tends to have a shorter incubation period. The variance of their incubation period also seems to be smaller. The Mann-Whitney rank test shows that there's a statistically significant difference between the incubation of age<40 and age>=40 groups with the null hypothesis: the medians of incubation period between two groups are the same. The p-values for corresponding alternative hypotheses: the age<40 group has a smaller incubation median is 0.00474. It suggests that the age<40 group has a shorter incubation period than the age>=40 group. It suggests that COVID-19 could have a faster distribution speed than H7N9, but the same spread speed as SARS and MERS in terms of their incubation periods [22] . We also investigate the incubation period of 12 family cases and 47 non-family cases in the dataset. The Mann-Whitney rank test shows that there are not significant differences between family patients and non-family patients in terms of the median of incubation period. Our studies indicate that incubation periods of the age>=40 years and age<40 years groups not only statistically significant but also linearly separable in machine learning. It will be more interesting to estimate different incubation time for them separately. Furthermore, different quarantine time should be applied to the age>=40 years and age<40 years groups for their different incubation periods. Our ongoing work is to collect more qualified data to extend our existing results and investigate incubation of COVID-19 for different groups besides comparing our incubation estimation with other studies [23] .
----- Title: Prevalence and clinical features of 2019 novel coronavirus disease (COVID-19) in the Fever Clinic of a teaching hospital in Beijing: a single-center, retrospective study, date: February 27, 2020 (19 of 38) ---- Pediatric patients were not included in our study. An incubation period was elicited from 12 patients (57.1%), ranging from 2 to 10 days with a median of 6.5 days. Consistent to previous reports, the incubation period of our cases was in a range of 2-10 days, with a median of 6.5 days [12] [13] [14] .
----- Title: Analysis of epidemiological characteristics of coronavirus 2019 infection and preventive measures in Shenzhen China: a heavy population city, date: March 03, 2020 (22 of 38) ---- Compared with the reports from Wuhan, Hubei [11] , the age of infected population in Shenzhen was younger and decreasing gradually, in which 33 patients were children. Taking account of the incubation period (mostly 3-7 days, with mean of 3.7 days) and the time between symptom onset and confirm of the diagnosis (6 day on average) [9, 12] , the peak of new confirmed cases coincided with the implementation of serial preventive strategy and measures,indicating these preventive strategies and measures were effective in preventing transmission of COVID-19 in Shenzhen.
----- Title: Transmission and clinical characteristics of coronavirus disease 2019 in 104 outside-Wuhan patients, China, date: March 06, 2020 (29 of 38) ---- Mean age was 43 (rang, 8-84) years (including 3 children) and 49 (47.12%) were male. We surveyed eight infected couples, a total of 3 infants were closely lived with their parents, but none of them was infected. Just 3 children were infected from their parents or relatives. These observations further demonstrated that infant and child are not so susceptible as adult, that is consistent with the previous reports 2,3,6,12 . The median incubation period was 6 (rang, 1-32) days, of 8 patients ranged from 18 to 32 days. The incubation period of 8 patients exceeded 14 days.. hospital-associated infections in Wuhan 2, 6 . The median incubation duration was 6 days, ranged from 1 to 32 days; 8 patients got more longer incubation duration (18, 19, 20, 21, 23, 24 , 24 and 32 days) that more than 14 days. The incubation duration ranged from 1 to 32 days with the median time of 6 days which was similar to the reported patients 13 . A recent report warned us the incubation duration may extend to 24 days 14 . We also found the incubation duration of 8 patients ranged from 18 to 32 days, indicating that it may exceed 14 days which reported with the initial infections 3 .
----- Title: Clinical Characteristics of 34 Children with Coronavirus Disease-2019 in the West of China: a Multiple-center Case Series, date: March 16, 2020 (32 of 38) ---- We describe the clinical and epidemiological characteristics of paediatric patients to provide valuable insight into early diagnosis of COVID-19 in children, as well as epidemic control policy making. • The epidemiological model in children was characterized with dominant family cluster transmission and extended incubation period, which should be taken into consideration in policy making for epidemic control. Admitted children with laboratory-confirmed 2019-nCoV . As mentioned in the literature review, the morbidity of COVID-19 was reported as 0.9% among children age 0 -14 [1] . So far, this is the largest case series to present the clinical and epidemiological characteristics in children with COVID-19, as well as the first study to analyze the clinical features in . The epidemiological features of paediatric patients indicated dynamic observation was necessary for suspected cases in children due to extended incubation period. The most common initial symptom, fever, was identified in 26 children (76.47%) in our study, however it was presented in only 43.8% of adults patients on admission [1] . Notwithstanding the relatively limited samples, our findings offer valuable insight into early diagnosis and epidemic control of COVID-19 in children. The median incubation period was 10.50 (7.75 - 25.25) days. The median incubation period was 10.50 (7.75 -25.25) days. • The epidemiological model in children was characterized with dominant family cluster transmission and extended incubation period, which should be taken into consideration in policy making for epidemic control. In addition, the median incubation period and disease course were analysed for epidemiological and clinical features of paediatric patients with COVID-19. Regarding the disease course, the median incubation period was 10.50 (7.75 -25.25) days. In accordance with the features of indirect contact exposure, the incubation period was 10.50 (7.75 -25.25) days in paediatric patients, while 4.00 (2.00 -7.00) days of incubation was revealed for patients in all age groups [1] . The epidemiological features of paediatric patients indicated dynamic observation was necessary for suspected cases in children due to extended incubation period.
----- Title: Investigation of three clusters of COVID-19 in Singapore: implications for surveillance and response measures, date: March 17, 2020 (35 of 38) ---- One case, a 6-month old male infant, was asymptomatic until one spike of fever 2 days into hospital admission. We reported the median (IQR) incubation period of SARS-CoV-2. The median incubation period of SARS-CoV-2 was 4 days (IQR 3–6). To answer these questions, we report data for the first three clusters of COVID-19 cases in Singapore, the epidemi ological and clinical investigations done to ascertain disease characteristics and exposure types, and summary statistics to characterise the incubation period of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the serial interval between trans mission pairs. We reported the median (IQR) incubation period, defined as the duration between estimated dates of infection and reported symptom onset, using R. We reported the serial interval range between transmission pairs in the household cluster. The incubation periods are plotted in figure 2 . The median incubation period was 4 days (IQR 3-6). Other study limitations include the small sample size used to ascertain the incubation period, because primary cases could not be identified with certainty. Based on symptomonset dates of 17 local cases, the median incubation period (4 days) corroborates other published findings.
----- Title: Early epidemiological analysis of the coronavirus disease 2019 outbreak based on crowdsourced data: a population-level observational study, date: January 01, 2020 (36 of 38) ---- Few patients (13 [3%]) were younger than 15 years and the age profile of Chinese patients adjusted for baseline demographics confirmed a deficit of infections among children. Adjustment for the age demographics of China confirmed a deficit of infections among children, with a RR below 0·5 in patients younger than 15 years (figure 1). We found a heavy skew of infection towards older age groups, with substantially fewer children infected. However, a substantial portion of the patients in our database are travellers, a population that is usually predominantly adults (although does not exclude children). Nevertheless, we would also expect children younger than 5 years to be at risk of severe outcomes and to be reported to the healthcare system, as is seen for other respiratory infections. A detailed analysis of one of the early COVID-19 clusters by Chan and colleagues 19 revealed symptomatic infections in five adult members of the same household, while a child in the same household aged 10 years was infected but remained asymptomatic, potentially indicating biological differences in the risk of clinical disease driven by age. Previous immunity from infection with a related coronavirus has been speculated to potentially protect children from SARS, 20, 21 and so might also have a role in COVID-19. Patient-level information is important to estimate key time-to-delay events (such as the incubation period and interval between symptom onset and visit to a hospital), analyse the age profile of infected patients, reconstruct epidemic curves by onset dates, and infer transmission parameters. The estimated incubation period in our data aligns with that of previous work. We estimated the duration of the incubation period on the basis of our line list data. The incubation period was estimated as the midpoint between the time spent in Wuhan and the date of symptom onset. On the basis of 33 patients with a travel history to Wuhan, we estimated the median incubation period for COVID-19 to be 4·5 days (IQR 3·0-5·5; appendix p 2). A useful feature of our crowdsourced database was the availability of travel histories for patients returning from Wuhan, which, along with dates of symptom onset, allowed for estimation of the incubation period here and in related work. Several teams have used our dataset and datasets from others to estimate a mean incubation period for COVID-19 to be 5-6 days (95% CI 2-11). [ 13] [14] [15] [16] The incubation period is a useful parameter to guide isolation and contact tracing; based on existing data, the disease status of a contact should be known with near certainty after a period of observation of 14 days.
----- Title: Characteristics of COVID-19 infection in Beijing, date: February 27, 2020 (37 of 38) ---- The median age of patients was 47.5 years (rang 6 months to 94 years;95% CI:45.1 to 49.9, Table 1 ); of them, 8 (3.1%) were children younger than 12 years old; 48 (18.3%) were 65 years age of older, 3 infants (two female, 6 months and 9 months respectively; a male 10 months) and a 25-year-old pregnant woman were infected, the gestational age was 33 weeks. The median incubation period was 6.7 days, the interval time from between illness onset and seeing a doctor was 4.5 days. The median time from contact symptomatic case to illness onset, which is called the incubation period, was 6.7 days, from illness onset to visit hospital was 4.5 days, from visit hospital to defined confirmed case was 2.1 days. 9 The median time of incubation period was 6.7 days.
----- Title: Epidemiological and clinical features of COVID-19 patients with and without pneumonia in Beijing, China, date: March 03, 2020 (38 of 38) ---- CD8+ T cell exhaustion might be critical in the development of COVID-19.. Before 2002, 4 kinds of coronaviruses (CoVs, namely HCoV 229E, NL63, OC43, and HKU1) were known to infect humans, causing 10%-30% mild upper respiratory infection in adults, occasionally severe pneumonia in elders, infants, and immunodeficient persons. Incubation period was calculated using the data of 31 cases with clear cutting time points of exposure and illness onset. The mean incubation period of COVID-19 was . A previous study carried out in Wuhan indicated that the mean incubation period was 5.2 days. 28 Finally, we characterized the epidemiological features including the incubation period, time to RT-PCR conversion of SARS-CoV-2, COVID-19 course, and the transmissibility SARS-CoV-2 in asymptomatic carriers.
Three articles show up of interest:
- (Han 2020) Estimate the incubation period of coronavirus 2019 (COVID-19)
- (Zhang et al 2020) Clinical Characteristics of 34 Children with Coronavirus Disease-2019 in the West of China: a Multiple-center Case Series
- (Henry and Oliveira 2020) Preliminary epidemiological analysis on children and adolescents with novel coronavirus disease 2019 outside Hubei Province, China: an observational study utilizing crowdsourced data
The first paper by (Han 2020) suggests that the incubation period is shorter for patients under the age of 40. The distribution of data points from Figure 3 of the paper appears to show a relatively short incubation period for those under 25. However there are only 59 patients in total for this study.
from PIL import Image
from matplotlib import pyplot as plt
image = Image.open("age_incubation.png")
fig, ax = plt.subplots(figsize=(18,9))
ax.imshow(image)
fig.suptitle("Figure 3: from (Han 2020) ", fontsize=18,weight='bold')
fig.subplots_adjust(top=1.1)
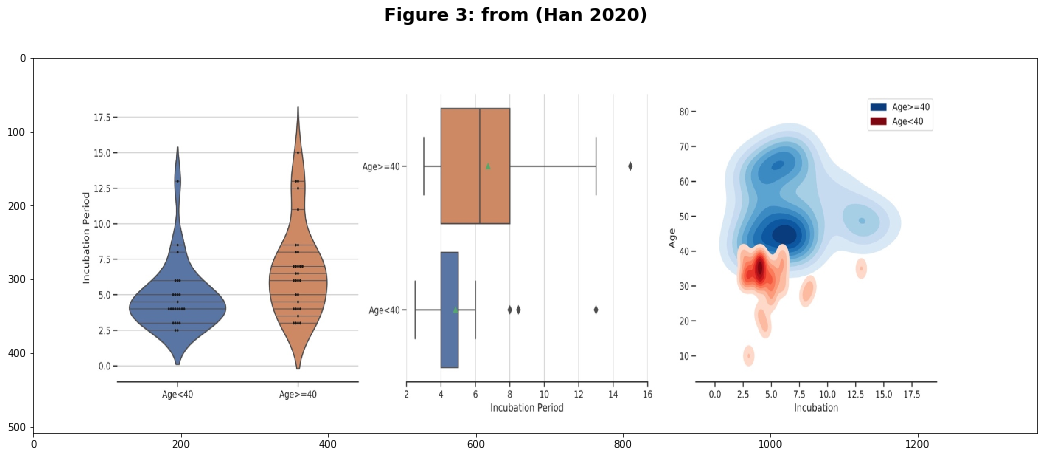
In (Zhang et al 2020) they suggest the opposite effect: the median incubation period of 10.5 days for pediatric patients, but only 4 for all age groups! However, this dataset also has a small sample size: 34. Lastly in (Henry and Oliveira 2020), the authors provide no new data to estimate the incubation period but instead reference (Cai et al 2020) which estimates a median incubation period of 6.5 days with n=10.
Unfortunately the point estimates for the incubation period in the general population appears to quite noisy. Furthermore, there is contradictory evidence about whether there is an age-based discrepancy in the average or median incubation period. Therefore the evidence from the CORD-19 corpus appears to be that incubation period averages around 5-6 days, and that there is not sufficient evidence to reject a difference in moments between a pediatric and adult population with regards to incubation time.